mne.read_evokeds#
- mne.read_evokeds(fname, condition=None, baseline=None, kind='average', proj=True, allow_maxshield=False, verbose=None)[source]#
Read evoked dataset(s).
- Parameters:
- fnamepath-like
The filename, which should end with
-ave.fifor-ave.fif.gz.- condition
intorstr|listofintorstr|None The index or list of indices of the evoked dataset to read. FIF files can contain multiple datasets. If None, all datasets are returned as a list.
- baseline
None|tupleof length 2 The time interval to consider as “baseline” when applying baseline correction. If
None, do not apply baseline correction. If a tuple(a, b), the interval is betweenaandb(in seconds), including the endpoints. IfaisNone, the beginning of the data is used; and ifbisNone, it is set to the end of the interval. If(None, None), the entire time interval is used.Note
The baseline
(a, b)includes both endpoints, i.e. all timepointstsuch thata <= t <= b.Correction is applied to each channel individually in the following way:
Calculate the mean signal of the baseline period.
Subtract this mean from the entire
Evoked.
If
None(default), do not apply baseline correction.Note
Note that if the read
Evokedobjects have already been baseline-corrected, the data retrieved from disk will always be baseline-corrected (in fact, only the baseline-corrected version of the data will be saved, so there is no way to undo this procedure). Only after the data has been loaded, a custom (additional) baseline correction may be optionally applied by passing a tuple here. PassingNonewill not remove an existing baseline correction, but merely omit the optional, additional baseline correction.- kind
str Either ‘average’ or ‘standard_error’, the type of data to read.
- projbool
If False, available projectors won’t be applied to the data.
- allow_maxshieldbool |
str(defaultFalse) If True, allow loading of data that has been recorded with internal active compensation (MaxShield). Data recorded with MaxShield should generally not be loaded directly, but should first be processed using SSS/tSSS to remove the compensation signals that may also affect brain activity. Can also be “yes” to load without eliciting a warning.
- verbosebool |
str|int|None Control verbosity of the logging output. If
None, use the default verbosity level. See the logging documentation andmne.verbose()for details. Should only be passed as a keyword argument.
- Returns:
See also
Notes
Changed in version 0.23: If the read
Evokedobjects had been baseline-corrected before saving, this will be reflected in theirbaselineattribute after reading.
Examples using mne.read_evokeds#
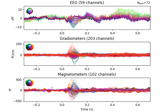
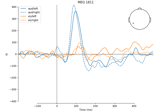
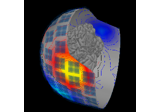
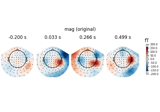
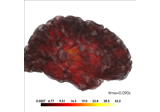
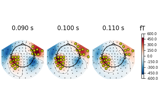
Source localization with equivalent current dipole (ECD) fit
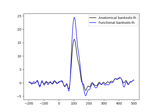
The role of dipole orientations in distributed source localization
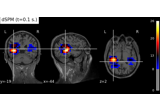
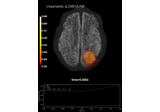
Compute MNE-dSPM inverse solution on evoked data in volume source space
Compute a sparse inverse solution using the Gamma-MAP empirical Bayesian method
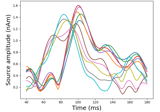
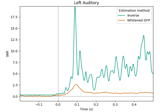
Extracting the time series of activations in a label

Compute sparse inverse solution with mixed norm: MxNE and irMxNE
Compute MNE inverse solution on evoked data with a mixed source space
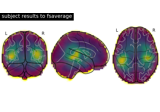
Plot point-spread functions (PSFs) and cross-talk functions (CTFs)
Compute spatial resolution metrics in source space
Compute spatial resolution metrics to compare MEG with EEG+MEG